# ProtFlow
**Repository Path**: william2015c/ProtFlow
## Basic Information
- **Project Name**: ProtFlow
- **Description**: No description available
- **Primary Language**: Unknown
- **License**: MIT
- **Default Branch**: master
- **Homepage**: None
- **GVP Project**: No
## Statistics
- **Stars**: 0
- **Forks**: 0
- **Created**: 2026-02-13
- **Last Updated**: 2026-02-13
## Categories & Tags
**Categories**: Uncategorized
**Tags**: None
## README
[](https://github.com/mabr3112/ProtFlow/actions/workflows/pytest.yaml)
# ProtFlow
A Python package for running protein design tools on SLURM clusters.
# Installation
First, clone the repository
```
git clone https://github.com/mabr3112/ProtFlow.git
```
ProtFlow requires python >= 3.11. You can either install it into an existing environment, or you can create a new one with like this:
```
conda create -n protflow python=3.11
conda activate protflow
```
Install the package in development mode (we are currently under active development).
```
cd ProtFlow
pip install -e .
```
After the install, instantiate your config.py file. This can be done with the command below, which will copy the config.py file from protflow/config_template.py
```
protflow-init-config
```
# The Configuration File
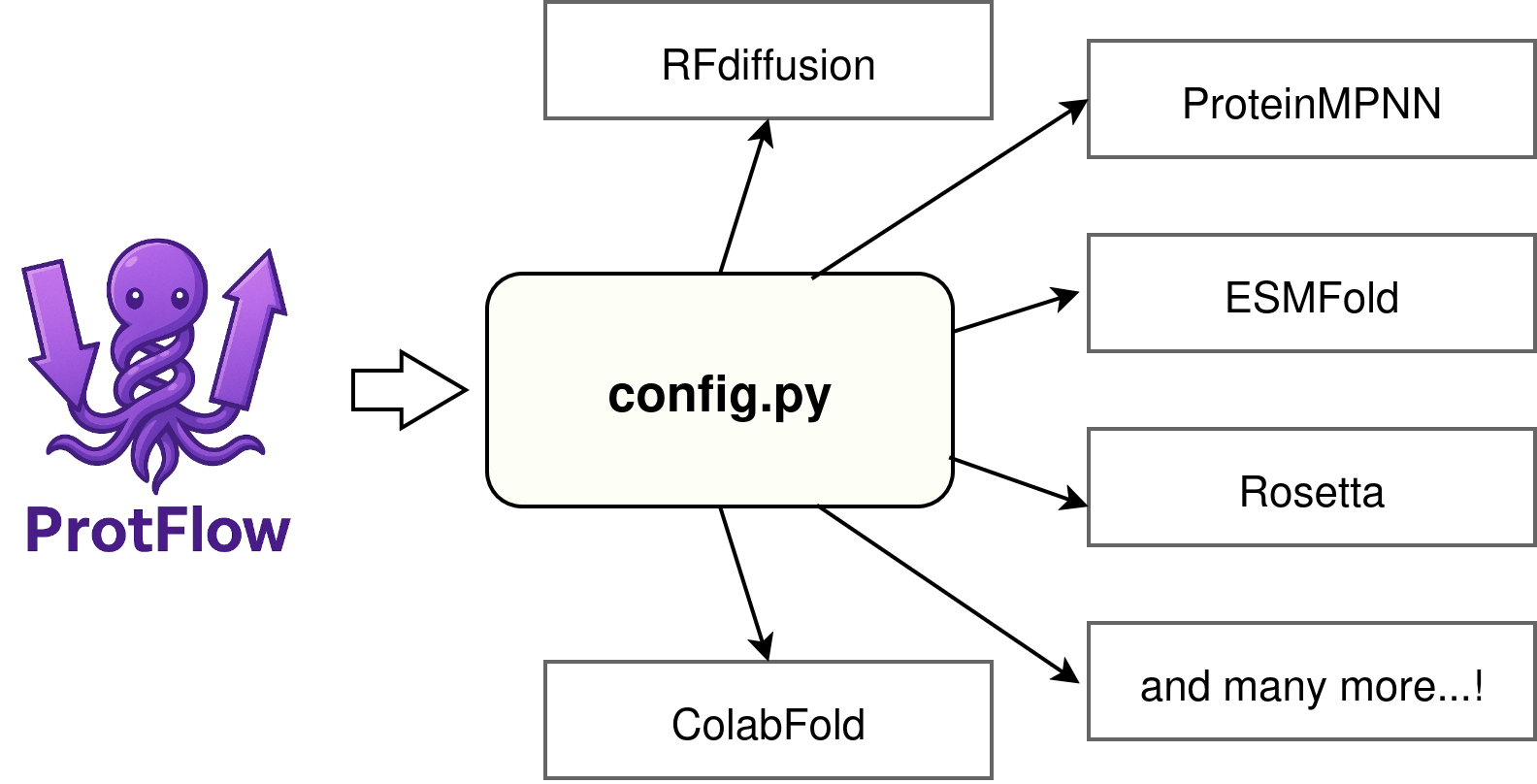
You will need to link protflow's runners to the individual python environments you have set up on your system. By Default, ProtFlow looks for environment paths in the file 'config.py' which we instantiated during the installation above.
If you try to initialize a runner for which you have not set paths in your config.py file, protflow will throw an error directing you to where you will need to set the required paths.
By default ``protflow-init-config`` will set up config.py in ~/.config/protflow/config.py.
If you want to link your protflow installation to a different config.py file elswhere, there is a simple cli-tool, ``protflow-set-config``.
Linking your protflow to a different config.py might come in handy if you want to share it system wide on a computing cluster with multiple users.
Simply provide the path to the target config.py to the tool:
```
protflow-set-config /path/to/your/config.py
```
If you want to unset this custom override, supply the --unset flag:
```
protflow-set-config --unset
```
Finally, to check which config.py file your protflow is currently using:
```
protflow-check-config
```
# Overview
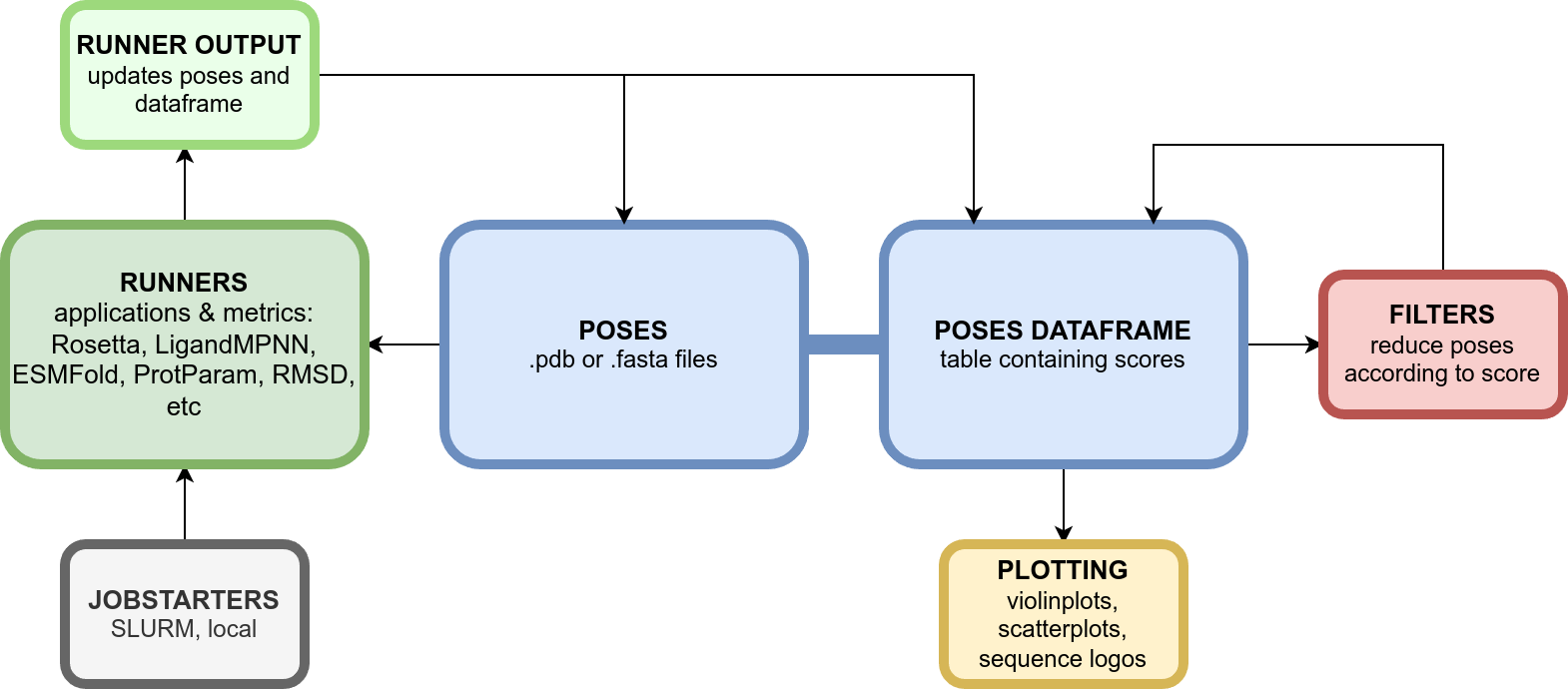
# Documentation
ProtFlow is documented with read-the-docs and sphinx. The documentation can be found here:
https://protflow.readthedocs.io/en/latest/

